About
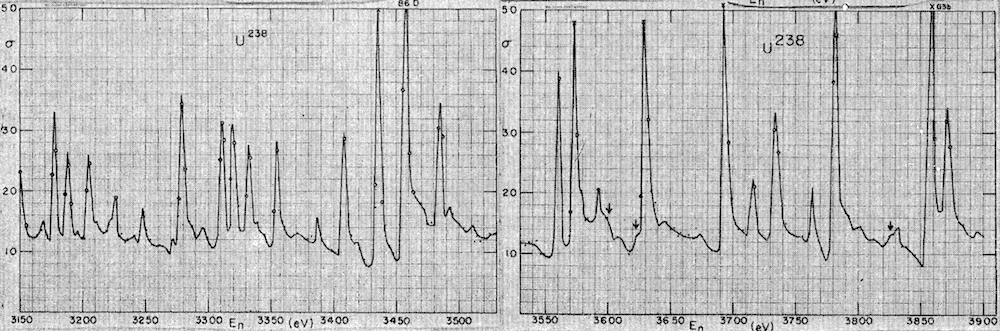
Random Matrix Theory is an area of mathematics that deals with matrix-valued random variables. There are many interesting questions that can asked for these mathematical objects, and this theory has found numerous applications in science over the years, such as biology, quantum physics, computer science... The study of eigenvalues of random matrices has many roots, including early work of Wishart in statistics, Wigner in nuclear physics, as well as Goldstein and von Neumann in numerical analysis.
This 1-day Symposium on Random Matrix Theory and its Applications, organized by Andrew Blumberg, Mathieu Carrière, Ivan Corwin and Raul Rabadan, is part of the events supported by the Columbia University Center for Topology of Cancer Evolution and Heterogeneity (Director Dr Rabadan), which is part of the National Cancer Institute's Physical Sciences in Oncology Network, and the "Probability and Society" initiative for Columbia. The purpose of the symposium is to present foundations and applications of Random Matrix Theory in biology and computer science.
For any questions or concerns please contact Mathieu Carrière at mc4660@cumc.columbia.edu or (917)-941-5182
Close
Speakers
Andrew Blumberg, PhD (University of Texas at Austin)
Luis Aparicio, PhD (Columbia University)
Ivan Corwin, PhD (Columbia University)
Ben Landon, PhD (MIT)
Alex Bloemendal, PhD (Broad Institute)
Jeff Pennington, PhD (Google Brain)
Jonathan Bloom, PhD (Broad Institute)
Close
Symposium Program
Friday November 1st:
9.30am - 10.00am --- Coffee/Tea & Registration
10.00am - 10.30am: Andrew Blumberg, PhD, University of Texas at Austin.
Homology of random point clouds
TBA
10.30am - 11.00am: Luis Aparicio, PhD, Columbia University.
Applications of random matrix theory to single-cell biology
11.00am - 11.30am --- Coffee break
11.30am - 12.00pm: Ivan Corwin, PhD, Columbia University.
Products of thin random matrices and random walks in random media
I will consider the asymptotic behavior of products of independent thin (e.g. tridiagonal) random matrices. Matrix entries in these products can be interpreted as partition functions for random polymer models, or as transition probabilities for random walks in random media. Through special "exactly solvable" models, we will explain a "Kardar-Parisi-Zhang universality" conjecture for the asymptotic behavior of these products. We will also explain how these systems can be thought of as toy models for certain biological phenomena.
Slides
12.00pm - 12.30pm: Alex Bloemendal, PhD, Broad Institute.
GWAS and BBP: Uncorrected confounding in genetic association studies
Genetic structure in human populations, i.e. systematic differences in allele frequencies between groups of different ancestries, can confound genome-wide association studies. The standard approach uses PCA to capture and correct for such structure; it can be inadequate in the face of the now well-known breakdown of PCA in the high-dimensional regime. We introduce and validate methods to characterize this problem and predict its extent in real genetic data.
Slides
12.30pm - 2.00pm --- Lunch break
2.00pm - 2.30pm: Ben Landon, PhD, MIT.
Extremal eigenvalues of sparse random matrices
Extremal eigenvalues of random matrices are of interest in statistical applications, and the random matrices that arise in such applications are may be sparse. We discuss the effect of sparsity on the asymptotic behavior of the extremal eigenvalues in some simple random matrix ensembles. Two such effects are higher order corrections to the spectral edges of Wigner's semicircle law, and a transition from Tracy-Widom to Gaussian fluctuations of the extremal eigenvalues.
Slides
2.30pm - 3.00pm: Jeffrey Pennington, PhD, Google Brain.
Operator-valued free probability meets deep learning: training and generalization dynamics in high dimensions
One of the distinguishing characteristics of modern deep learning systems is that they typically employ neural network architectures that utilize enormous numbers of parameters, often in the millions and sometimes even in the billions. While this paradigm has recently inspired a broad research effort on the properties of large networks, relatively little work has been devoted to the fact that these networks are often used to model large complex datasets, which may themselves contain millions or even billions of constraints. In this talk, I will present a formalism based on operator-valued free probability that enables exact predictions of training and generalization performance in the high-dimensional regime in which both the dataset size and the number of features tend to infinity. The analysis provides one of the first analytically tractable models that captures the effects of early stopping, over/under-parameterization, explicit regularization, and which exhibits the characteristic double-descent curve.
Slides
3.00pm - 3.30pm --- Coffee break
3.30pm - 4.00pm: Jonathan Bloom, PhD, Broad Institute.
Loss landscapes, Morse theory, and linear autoencoders
Random matrix theory has guided intuition on loss landscapes of deep neural networks, though rigorous connections invoke unrealistic assumptions. I’ll describe a simple model, the linear autoencoder, where the critical values are sums of eigenvalues of the data and the dynamics of learning may be understood through the topological lens of Morse theory. We recently extended this analysis to L2-regularized linear autoencoders, proving all critical points are symmetric, with implications for PCA algorithms and the biological plausibility of backprop. I'll speculate that Morse theory may give intuition complementary to RMT in deep models as well. Based on “Loss Landscapes of Regularized Linear Autoencoders” (ICML 2019) with Daniel Kunin, Aleksandrina Goeva, and Cotton Seed at the Broad Institute.
Close
Location
Columbia University
Lerner Hall 477
2920 Broadway,
New York, NY 10027
Close



